Untangling the Osquery❓ tables web🕸 using Jupyter Notebooks📓 | Part 1
In this blog I want to take you on the journey of discovery I started not so long ago.
My journey started when I came across ThreatHunter-Playbook from Roberto Rodriguez and Jose Luis Rodriguez. In this GitHub repository they talk about Jupyter Notebooks, at first I did not understand it at all. 🤔
I needed some time to digest all this new knowledge. 📚
The Jupyter Notebook is an open-source web application that allows you to create and share documents that contain live code, equations, visualizations and narrative text. Uses include: data cleaning and transformation, numerical simulation, statistical modeling, data visualization, machine learning, and much more.
As time went by I started to understand the genius of it and how you might be able to apply it in Information Security and Threat Hunting.
I come from a System and Network Engineering background before rolling into Information Security. Let’s say programming/development is not my forte.
So I had to take quite some extra time learning Python and how it is applied in Jupyter Notebooks, I am still learning new things everyday… 😅
But the best way to learn something new is to dive right in, so since I discovered Jupyter Notebooks I have been using it in pretty much anything I need to put some programmatic logic in. For example: CTFs, API calls, Machine Learning and Threat Hunts.
One of the latest Jupyter Notebook projects I have been working on is the subject of this blog.
I really like the current Open Source Security community and one special project for me is Osquery.
Osquery exposes an operating system as a high-performance relational database.
This gives the possibility to “question” a system with simple SQL-like query syntax:
Find Jupyter Notebook processes running at this moment.Query
SELECT name,path,cmdline FROM processes WHERE cmdline LIKE '%jupyter%';Output
name = python.exe
path = C:\Users\username\Anaconda3\python.exe
cmdline = C:\Users\username\Anaconda3\python.exe C:\Users\username\Anaconda3\Scripts\jupyter-notebook-script.py C:\Users\username/
Osquery is a great tool, as you can query almost everything on a system real-time, all is built around the simple concept of making difficult system calls easy to question (query) in SQL-like syntax. Every component of the system that you can query is a different table, as you can understand there are quite some tables. At the moment of writing Osquery is at version 4.3.0 and there are 257 tables! 😲
Not all tables can be used on all Operating Systems, at this moment it is divided as follows:
- Windows: 88 tables
- Linux: 149 tables
- macOS: 184 tables
- FreeBSD: 48 tables
There is a great amount of information that you can retrieve from different Osquery tables, so in order to find related context for my Threat Hunts, I had to manually search through the table structures and sometime with trial and error find the data I needed.
These manual tasks made me wonder…
How can I visualize the relationship, for possible JOINs, between the tables?
Visualization of the connection between tables can help to understand the possibilities and how the tables can be extended.
Here is where my “Osquery❓ tables ➕ Jupyter Notebook📓” journey started.
I was working on this as a personal project but when I heard about Jupyterthon I thought it was a good opportunity to present my research and also write this blog.
To understand the code snippets in the Jupyter Notebook you might need some basic understanding of:
My complete Jupyter Notebook with all the steps are available in my GitHub repository, so I will not go into full detail of every step. I will only highlight the most interesting sections in this blog.
When I start a Jupyter Notebook project I always try to understand the basic data I have.
First I had to understand how I could get the data I wanted in the Jupyter Notebook.
I looked at the repository structure of the Osquery GitHub to help me understand how I could extract the table information to make it usable.
Side-note: Not a developer at all, code could have been probably much better. Most of my time went into translating my thoughts into Python with quite some help from Google. 😅
The table definitions of all Osquery tables is well-documented and structured so extracting them and putting them in DataFrames was not that difficult.
DataFrames can be used to store and manipulate tabular data in rows of observations and columns of variables. You can see it as some kind of Excel spreadsheet.
I had to do a bit of hacking to make the Table row length the same as the Column rows, as every Table has more than one Column.
After this I did some matching against the CMakelists.txt to get the correct association based on OS, I also had to do some manual associations because of a few parsing issues, but all those juicy details can be read in my Jupyter Notebook. 😉
Now let’s get to the reason why I started this journey in the first place, some graphs!
Graphs give you visual representation of relationships, I use it to visualize the relationship between Osquery tables and columns. By studying the graphs, you can find how you could use different tables to extract data and you also could find unexpected relationships.
Just for fun I created a graph without any kind of filtering. 🎉😁

Beautiful chaos right? … 🕸 📈
This graph is interactive if you click here you will be able to see the moving graph in all its glory, you can zoom in and see the name of each node. Each node represents a table or a column.
In the graph I used 3 different color nodes:
- Orange are for the tables with only 1 connection
- Green represent the tables with more than 1 connection
- Red are all the columns that are used to connect the tables to each other and the size is dependent on how many connections there are.
If you were wondering, this beautiful interactive graph rendering comes from a combination of the following Python packages.
You can very easily import new packages in your Jupyter Notebook, if the packages are not installed, you can download them and use them immediately in your Jupyter Notebook. Below is how I downloaded the graph packages.
The assumptions I made to create the graph were that:
- If column name has only one connection (based on
nx.degree()) discard, this would mean the column only exists in that table. - If there are tables that do not have a connection because of the removal above also discard them.
- Connect tables based on the same column name.
Of course I know you can JOIN based on other column names and make complex queries, but the graphs can be a starting point to give an idea of the possible connections.
Next step was to filter it down based on OS. Even after the OS filtering there was still too much chaos, as there are many column names that are too generic to be able to use them for any kind of meaningful connection. I started trimming it down first by looking at the most common column names.
After filtering out the most obvious generic column names like name, description, type, size etc. I went through the graph, node by node, removing column names that did not have the same meaning in the different tables, this helped quite a bit in reducing the noise.
The filtering applied is a bit broad but later it can be fine-tuned, this will be done in the next release.
For now this filtering will suffice to illustrate the power of graphing the Osquery tables.
After doing the above filtering I ended up with the following Windows graph.
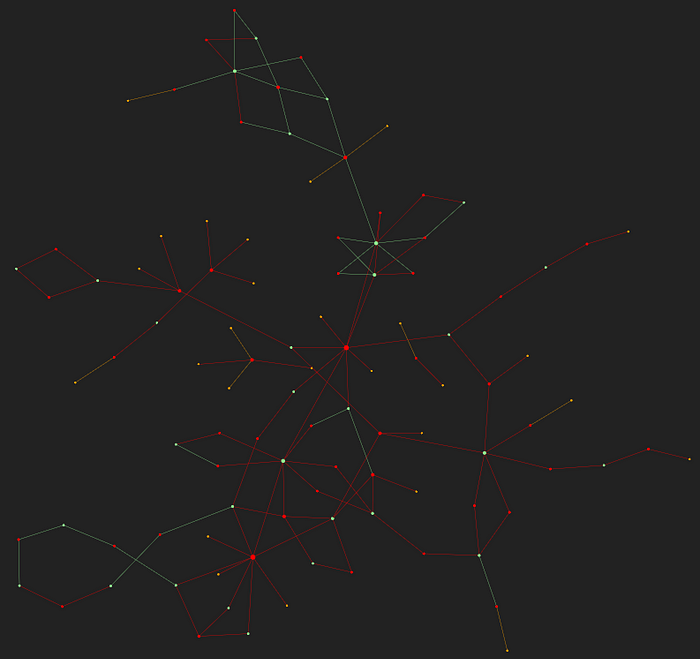

It looks like some kind of constellation. 🎇🤣
Click here for the full interactive graph.
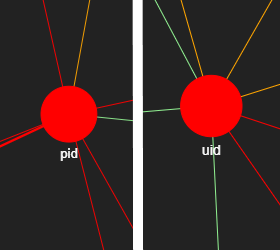
Zooming in on the graph you notice that uid and pid are the most connected column names. This would mean if a table has one of these columns it will give the most possibility for extra enrichment based on other tables.
It is also good to be able to check if the graph gives results that you would expect.
Like listening_ports and process_open_sockets tables, they are both based on similar data so it would be logical that the column names would be pretty much the same.
Interesting to see if you analyze the graph below you see almost all columns are the same but port and address(not visible), I went back to the table schema and noticed a difference in naming. In process_open_sockets the port and address have a prefix, local_ and remote_.
This would mean if the listening_ports columns are renamed with the local_ prefix (as listening is local), it will make it more alike. I might send in a Feature Request for this. There is one more column that is the same but this was removed by the filtering, the name was too generic.

Another part of the project was to be able to answer the question…
How can I get from table A to table B?
This can help during investigations to quickly pivot to other tables and can also expose paths that are not obvious at first.
To do this I created a function to calculate the shortest path based on Dijkstra’s algorithm. Usually you also need to add “weight” to the lines/edges in the graph, but this is for more complex graphs.
I just wanted to visualize the shortest path possible between the tables, this sometimes gives unexpected results. By looking at the shortest path you can possibly write shorter queries bypassing the most obvious JOIN tables, you could also investigate column names that might have incorrect name or values.
As I wanted some interactivity in the Jupyter Notebook to choose which table is A and which is B I added interactivity by adding from ipywidgets import interact module.

Above graph gives an interesting connection, in theory this would mean that you can JOIN a PowerShell script execution from the powershell_events¹ table to the windows_events² table based on datetime.
And using a time-range based on that datetime you can display the Windows events that happened shortly before or after the PowerShell script execution.
Side-note: Not a SQL expert so I am not sure if it will work like this.
¹ Table holds executed PowerShell scripts.
² Table holds Windows Event logs.
You can play with it for yourself using my Jupyter Notebook, it is interesting to go through the tables and look at the different paths.
For now, I only focused on the Windows graph, Linux and macOS graphs need a bit more filtering as they have more or less double the amount of tables compared to Windows.
Graphs for Linux, macOS and FreeBSD will follow later.
Now that you reached the end of my blog, you might be wondering what are the other applications besides just viewing colorful graphs.
- One application is using it for Incident Response or Threat Hunting, an example, you run a query and get information that need further investigation, if you can easily find out which tables you can use, to pivot to, based on the results of the first query, you can enrich your results faster without having to look at all the tables and with trial and error.
- Another application is understanding the relationship between tables by analyzing the graphs and possibly find hidden paths.
- Visually examine tables and columns, you can find for example discrepancies or column names that can be renamed to consolidate naming.
I would also like to share some findings based on this research:
- It is quite difficult to create connections based only on column name, as some columns have the same name but different kind of data. It might be an idea to add some kind of data identifier to the column name, that way you always know what kind of data to expect and possibly use it for JOIN definitions.
- Naming convention might also help, as the amount of tables grow it might get more difficult to see the relationships between tables. Some tables have different names or a permutation of the same name, but the fields have the same data.
While this is a good start there is still some work to do, I will be working on following next:
- Create graphs for Linux, macOS and FreeBSD.
- Check the data returned from all the tables when querying and use that data to further fine-tune the filtering.
- Not all shortest paths are correct, data in the columns do not match, maybe use the same data as previous bullet to build better paths.
I would like to thank my colleagues for their time and also Roberto Rodriguez and Josh Brower for their feedback and improvements.
